March 22, 2013, 7:15 p.m. by Rosalind Team
Topics: Combinatorics, Dynamic Programming, String Algorithms
Breaking the Bonds
In “Perfect Matchings and RNA Secondary Structures”, we considered a problem that required us to assume that every possible nucleotide is involved in base pairing to induce an RNA secondary structure. Yet the only way this could occur is if the frequency of adenine in our RNA strand is equal to the frequency of uracil and if the same holds for guanine and cytosine.
We will therefore begin to explore ways of counting secondary structures in which this condition is not required. A more general combinatorial problem will ask instead for the total number of secondary structures of a strand having a maximum possible number of base pairs.
The graph theoretical analogue of the quandary stated in the introduction above
is that if we have an RNA string
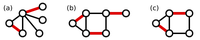
In light of this fact, we define a maximum matching in a graph as a matching containing as many edges as possible. See Figure 2 for three maximum matchings in graphs.
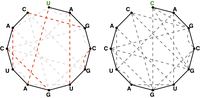
A maximum matching of basepair edges will correspond to a way of forming as many base pairs as possible in an RNA string, as shown in Figure 3.
Given: An RNA string
Return: The total possible number of maximum matchings of basepair edges in the bonding graph of
>Rosalind_92 AUGCUUC
6