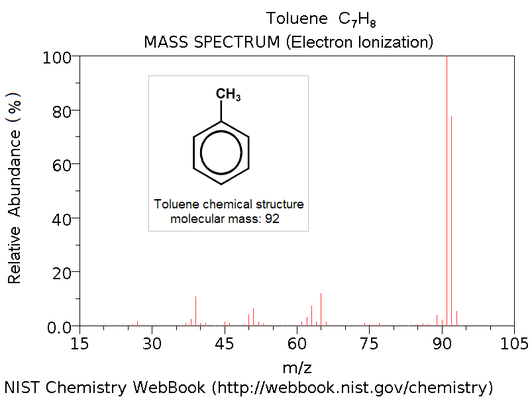
A mass spectrum is the output of a mass spectrometer, which vaporizes an unknown sample (often a peptide) and ionizes the resulting particles. Then, the spectrum is created by plotting ions' possible mass-charge ratios against the actual observed frequency of these ions in the sample. This creates a graph with different "peaks"; the goal is to reconstruct chemical properties of the sample (e.g., the amino acid sequence of a peptide) from the mass-charge plot.
On the Rosalind site, we will use the term "mass spectrum" to refer to a simpler collection of all fragment masses obtained from the sample. For example, if we have repeated peptide fragment masses 199, 178, and 315, then we need to use a table containing all possible amino acid masses (like the average mass table) and infer that the fragments must have been "QA", "AT", and "ATH", leading us to reconstruct the short peptide "QATH".