Mass spectrometry is a method used to identify the chemical makeup of molecules by splitting these molecules into many small pieces and then analyzing these snippets.
Most commonly, mass spectrometry is applied to peptide sequencing. In a simple form of the problem that we will study, many copies of the same peptide are blasted into bits, and then these smaller pieces are weighed. Using these experimental masses and a table giving the mass of each amino acid used in peptide construction (like the average mass table), the goal is to reconstruct the unknown sequence of amino acids making up the peptide.
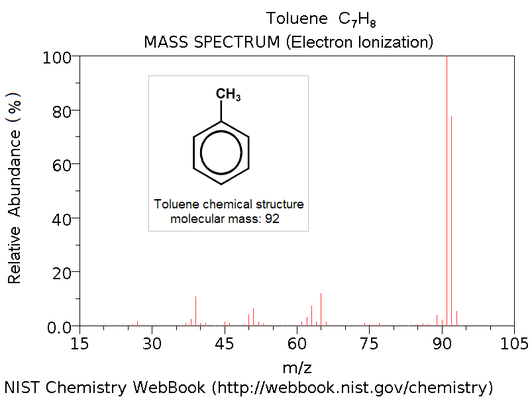
In practice, the unknown peptide fragments are ionized, and then the frequency of particles is measured by the mass spectrometer according to their mass/charge ratio. This provides us with a 2-dimensional plot of mass/charge to frequency as shown in the figure below; the computational problem is to use the distribution of "peaks" in this graph to infer chemical properties of the original peptide molecule, most notably its amino acid sequence.