Jan. 4, 2013, 5:48 p.m. by Rosalind Team
Topics: Graph Algorithms, String Algorithms
If At First You Don't Succeed...
We introduced the problem of finding a motif in a genetic string in “Finding a Motif in DNA”. More commonly, we will have a collection of motifs that we may wish to find in a larger string, for example when searching a genome for a collection of known genes.
This application sets up the algorithmic problem of pattern matching, in which we are searching a large string (called a text) for instances of a collection of smaller strings, called patterns. For the moment, we will focus on requiring that all matches should be exact.
The most obvious method for finding exact patterns in a text is to simply apply a simple "sliding window" algorithm for each pattern. However, this method is time-consuming if we have a large number of patterns to consider (which will often be the case when dealing with a database of genes). It would be better if instead of traversing the genome for every pattern, we could somehow only traverse it once. To this end, we will need a data structure that can efficiently organize a collection of patterns.
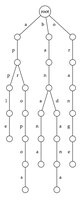
Given a collection of strings, their trie (often pronounced "try" to avoid ambiguity with the general term tree) is a rooted tree formed as follows. For every unique first symbol in the strings, an edge is formed connecting the root to a new vertex. This symbol is then used to label the edge.
We may then iterate the process by moving down one level as follows. Say that an edge
connecting the root to a node
As a result of this method of construction, the symbols along the edges of any path in the trie from the root to a leaf will spell out a unique string from the collection, as long as no string is a prefix of another in the collection (this would cause the first string to be encoded as a path terminating at an internal node).
Given: A list of at most 100 DNA strings of length at most 100 bp, none of which is a prefix of another.
Return: The adjacency list corresponding to the trie
ATAGA ATC GAT
1 2 A 2 3 T 3 4 A 4 5 G 5 6 A 3 7 C 1 8 G 8 9 A 9 10 T