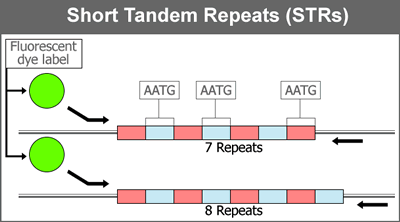
A microsatellite, or "short tandem repeat" (STR), is a 2-6 bp interval of DNA that is repeated several successive times in a genome; for example, "CA" occurs five times in the string "CACACACACA". When this region is copied by DNA polymerase during replication, a mistake can be made that adds or deletes a microsatellite occurrence, which creates an allele at that location of the genome. Accordingly, microsatellites can be used as markers in genetic fingerprinting.
Each individual has two copies of any microsatellite segment (one from each parent), and each of these copies can vary in the number of repeats, representing a particular allele. The figure below represents an example of parental testing using a single microsatellite.
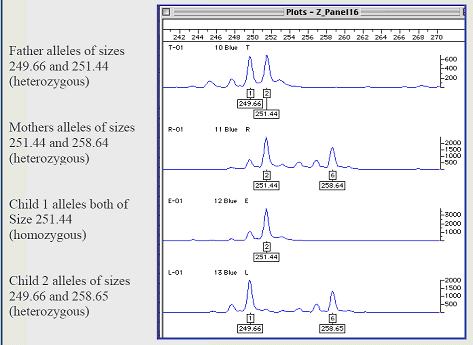
Microsatellites are analyzed by amplifying them in a polymerase chain reaction that uses specific primers to the conservative regions flanking the repeated region. As a simple example, if one individual has "CACACACACA" (five microsatellites) and another individual has "CACACACA" (four microsatellites), then amplifying them by a factor of 1 million will produce 10 million bases for the first individual and 8 million bases in the second individual, a substantial (and measurable) discrepancy.