Jan. 11, 2013, 6:26 p.m. by Rosalind Team
Topics: Set Theory
Genetic Fingerprinting
Recall that a restriction enzyme cuts the endpoints of a specific interval of DNA, which must form a reverse palindrome that typically has length 4 or 6. The interval of DNA cleaved by a given restriction enzyme is called its recognition sequence.
A single human chromosome is so long that a given recognition sequence will occur frequently throughout the chromosome (recall from “Expected Number of Restriction Sites” that a recognition sequence would be expected to occur several times even in a short chromosome). Nevertheless, the small-scale mutations that create diversity in the human genome (chiefly SNPs) will cause each human to have a different collection of recognition sequences for a given restriction enzyme.
Genetic fingerprinting is the term applied to the general process of forming a limited picture of a person's genetic makeup (which was traditionally cheaper than sequencing). The earliest application of genetic fingerprinting inexpensive enough to be widely used in common applications, like forensics and paternity tests, relied on a process called restriction digest. In this technique, a sample of DNA is replicated artificially, then treated with a given restriction enzyme; when the enzyme cuts the DNA at restriction sites, it forms a number of fragments. A second process called gel electrophoresis then separates these fragments along a membrane based on their size, with larger pieces tending toward one end and smaller pieces tending toward the other. When the membrane is stained or viewed with an X-ray machine, the fragments create a distinct banding pattern, which typically differs for any two individuals.
These intervals can be thought of simply as the collection of distances between restriction sites in the genome. Before the rapid advances of genome sequencing, biologists wanted to know if they could use only these distances to reconstruct the actual locations of restriction sites in the genome, forming a restriction map. Restriction maps were desired in the years before the advent of sequencing, when any information at all about genomic makeup was highly coveted. The application of forming a restriction map from cleaved restriction fragments motivates the following problem.
For a set
If
In practical terms, we can easily obtain a multiset
Given: A multiset
Return: A set
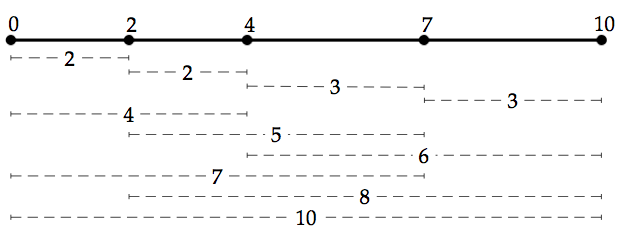
2 2 3 3 4 5 6 7 8 10
0 2 4 7 10